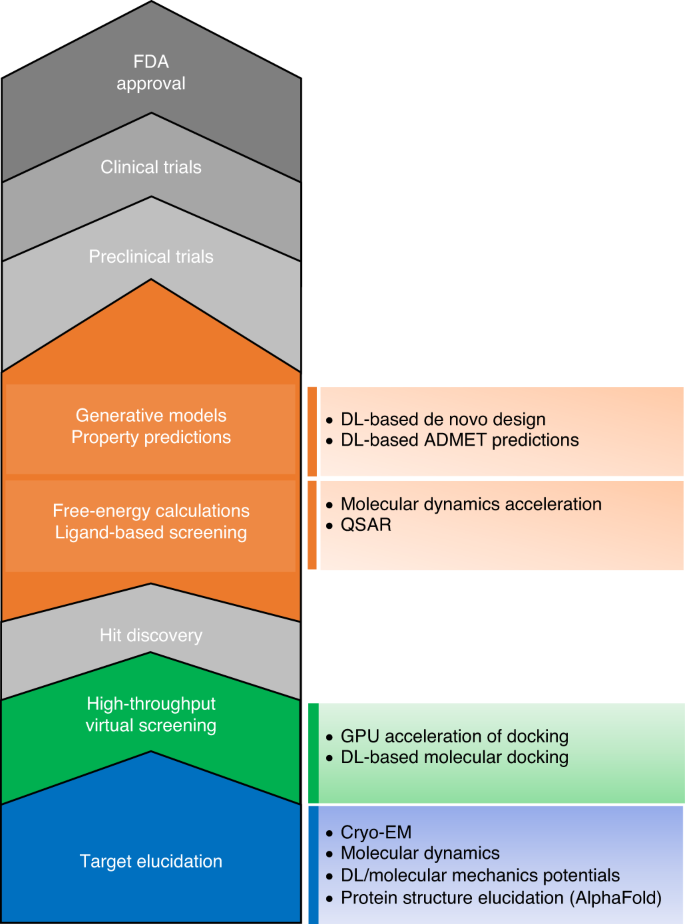
The transformational role of GPU computing and deep learning in drug discovery | Nature Machine Intelligence

Heterogeneous CPU+GPU-Enabled Simulations for DFTB Molecular Dynamics of Large Chemical and Biological Systems | Journal of Chemical Theory and Computation
GPU-Accelerated Molecular Dynamics Simulation to Study Liquid Crystal Phase Transition Using Coarse-Grained Gay-Berne Anisotropic Potential | PLOS ONE